Our research focuses on understanding (a) the genetic basis of between-individual variation in complex traits and (b) how such complex genetic architectures, instead of being static properties of a trait, get re-shaped when populations are exposed to different environments (genotype-by-environment interactions or GxE). In addition, we are particularly interested in (c) understanding how phenotypic robustness is regulated in such traits. That is, we aim to understand not only why individuals in a population look different from each other, but also why some individuals are more vulnerable than others when exposed to perturbations like stressful or new environments.
To explore these questions within the context of natural genetic variation, we use wild-derived outbred Drosophila melanogaster populations as model system. We integrate experimental and analytical tools across the fields of quantitative and population genetics, molecular and computational biology, and use experimental evolution to generate and analyze large-scale genomics datasets.
Conceptually, our research tackles long-standing questions in evolutionary biology including the genotype-phenotype map and its context-dependent nature, and the (apparent) conflict between robustness and evolvability.
***
Genetic Basis of Complex Traits
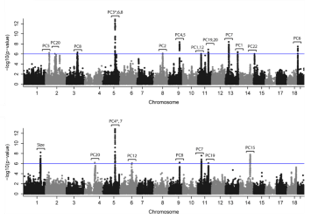
Identifying the genomic loci regulating between-individual variation at the population level is the first step to understand how information stored in the genome is translated into phenotypic variation. From transcriptional to morphological variation, we aim to understand the genetic architecture of such traits as the first step in the reconstruction of the genotype-phenotype map.
***
Genotype-by-environment interactions
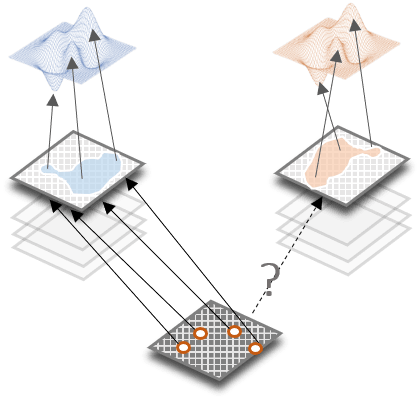
The genetic regulation of phenotypic variation is context-dependent and therefore individual genetic effects, as well as the structure of gene networks, get remodeled depending on environmental conditions. We focus on quantifying the contribution of genotype-by-environment interactions (GxE) to the regulation of complex trait variation with a particular focus on identifying cryptic genetic variation.
***
Genetic regulation of phenotypic robustness
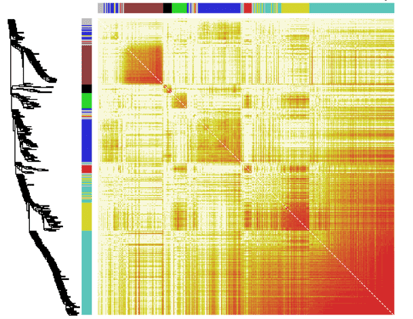
Phenotypic robustness represents the degree to which a trait is perturbed by genetic or environmental conditions. It can be considered as any other complex trait since it varies between individuals – some are able to withstand stress better than others. We focus on identifying the genetic basis of phenotypic robustness, that is, which genomic loci regulate the amount of noise (i.e., variance around the mean value of the trait) in morphological and transcriptional traits.
***
Experimental evolution to understand the adaptation process
To understand the trajectory of a population towards a new fitness peak, we use experimental evolution and follow, from a genetic and phenotypic perspective, the trajectory of adaptation to the new peak. With this approach we integrate the other research themes in the lab: Which loci are involved in the long-term adaptation process? Are they the same loci that respond to short-term exposure to new environments (i.e., are GxE loci regulating plasticity, adaptive?) What role does cryptic genetic variation play in adaptation? What is the role of ‘robustness loci’ in allowing populations to explore new phenotypes? How is phenotypic variance shaped by adaptation?

***